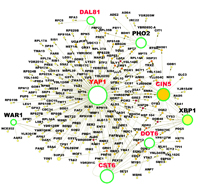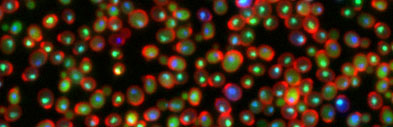
Global reorganization of the yeast proteome
in response to alkylation damage
SAMSONLAB MIT
| |
| GFPlibrary |
| downloads filesanalysis programs |
| single cell measurments |
| contact us |
| Welcome |
This website provides a means to reqeust the data files associated with the study by Mazumder et al (Nucleic Acids Research, 2013) describing the changes that occur in the proteome both in terms of levels and broad nuclear-to-cytoplasmic localization in response to the alkylating agent methyl methanesulfonate (MMS). The yeast GFP library (Huh et al, Nature, 2003; http://www.ncbi.nlm.nih.gov/pubmed/14562095) was used to investigate the response of >4000 individual GFP-tagged strains to MMS-induced genotoxis stress. Once responders had been identified, network analysis in terms of physical, genetic and transcription factors interactions, and Gene Ontology analysis, revealed the global cellular processes affected by MMS. Please see our paper for greater details of this study. Contact us to request access to the orginal images, the analysis programs and single cell files. The total size of the images is 2 TB. Hence it has been broken up into smaller units and compressed to facilitate easy transfer. The Excel file describes the distributions of the strains in the plates. Please contact "be-it at mit dot edu" should you have any questions or comments. Also follow this link to look at some previous studies from the Samson laboratory: http://genomicphenotyping.mit.edu
|
| Cellomics Arrayscanner | ||
|
||

